Why is Chromosome 17q21 linked to Asthma?
Sudipta Das and David H Broide
DOI10.21767/2471-304X.100016
Sudipta Das and David H Broide*
Department of Medicine, University of California, San Diego, La Jolla, USA
- *Corresponding Author:
- David H Broide
Department of Medicine
University of California San Diego
Biomedical Sciences Building
La Jolla, USA
Tel: +8585342374
E-mail: dbroide@ucsd.edu
Received date: June 10, 2016; Accepted date: August 18, 2016; Published date: August 20, 2016
Citation: Das S, Broide DH. Why is Chromosome 17q21 linked to Asthma? Insights Allergy Asthma Bronchitis. 2016, 2:2.
Abstract
Asthma is an inflammatory disorder of the airways associated with increased airway responsiveness and airway remodeling. Several genome wide association studies (GWAS) and non-GWAS studies have demonstrated a strong genetic linkage of chromosome 17q21 genes with asthma in populations of diverse ethnic backgrounds. In particular, SNPs in the 17q21 region are found within a large linkage disequilibrium (LD) block that contains not only the orosomucoid like-3 (ORMDL3) gene, but also includes other 17q21 genes i.e. gasdermin B (GSDMB), IKAROS family zinc finger 3 (IKZF3), and zona pellucida binding protein 2 (ZPBP2). Studies of mice expressing increased levels of the human ORMDL3 gene have demonstrated that these mice have spontaneous increased airway remodeling and airway responsiveness in the absence of airway inflammation. Further functional studies are needed to investigate whether some or all of these four genes could contribute to the pathogenesis of asthma.
Keywords
Asthma; Chromosome
Introduction
ORMDL3 is an allergen-inducible endoplasmic reticulum (ER) gene expressed in airway epithelial cells. ORMDL3 activates ATF-6α pathway during ER unfolded response and binds to sarco/endoplasmic reticulum Ca2+ ATPases pump (SERCA2b) to regulate ER-mediated Ca2+ signaling and cellular stress response [12,13]. ORMDL3 acts as a mediator of sphingolipid homeostasis [14,15] and can also regulate eosinophil trafficking, recruitment and degranulation [16]. Studies of mice generated to express increased levels of human ORMDL3 have also demonstrated that in vivo ORMDL3 activates ATF6α (but not Ire1 or PERK), and that this is associated with increased expression of SERCA2b. SERCA2b regulates calcium flux, protein synthesis, gene expression, and metabolism and has been implicated in airway remodeling in asthma. In addition, ORMDL3 transgenic mice have increased airway remodeling and airway responsiveness without increased airway inflammation [13]. Thus, there is biological evidence that ORMDL3 may be one of the genes on chromosome 17q21 contributing to the pathogenesis of asthma.
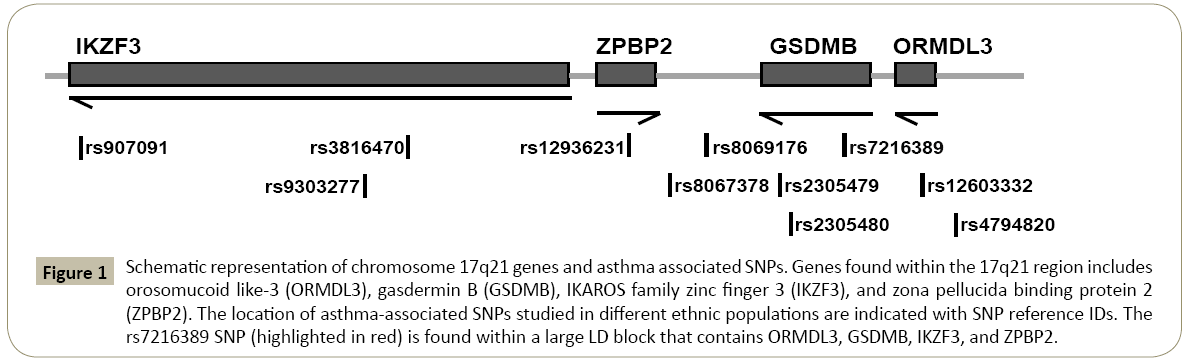
Figure 1: Schematic representation of chromosome 17q21 genes and asthma associated SNPs. Genes found within the 17q21 region includes orosomucoid like-3 (ORMDL3), gasdermin B (GSDMB), IKAROS family zinc finger 3 (IKZF3), and zona pellucida binding protein 2 (ZPBP2). The location of asthma-associated SNPs studied in different ethnic populations are indicated with SNP reference IDs. The rs7216389 SNP (highlighted in red) is found within a large LD block that contains ORMDL3, GSDMB, IKZF3, and ZPBP2.
Several studies show that asthma-associated variants of 17q21 alter the transcription levels of ORMDL3 and GSDMB. While several genetic studies have clearly demonstrated that ORMDL3 and GSDMB either individually, or in combination act as associated risk factors for asthma, no functional studies till date have established the biological role(s) of GSDMB in mediating asthma pathogenesis. GSDMB (also known as GSDML) is a novel protein that belongs to the gasdermin (GSDM) family that consists of four human genes: GSDMA, GSDMB, GSDMC and GSDMD [17]. A mouse or rat ortholog of the GSDMB gene has not been identified, and it may have evolved independently in humans through a GSDM gene duplication [18]. The human GSDMB gene consists of 12 exons and four different splice variants have been described, which differ in exons 6 and 7 of the GSDMB gene [19]. GSDMB has been linked to cancer progression and is expressed in human gastric, liver, colon and breast cancer cell lines and carcinomas [17,19]. Although several genetic studies show a strong association of GSDMB with asthma, no functional studies till date have investigated the biological role of GSDMB in asthma and related airway cell types. Interestingly, the rs7216389 SNP highly associated with asthma is localized within the GSDMB gene and demonstrates a strong LD with other 17q21 genes, thus underscoring the need to perform future studies on GSDMB in asthma.
The 17q21 region contains two other genes frequently associated with asthma, i.e., IKZF3 and ZPBP2. IKZF3 (Aiolos) belongs to the Ikaros family of zinc-finger proteins which encodes hematopoietic-specific transcription factors involved in the regulation of lymphocyte development [20]. IKZF3 plays a critical role in regulating B and T cell development, and several alternative coding as well as non-coding transcripts variants of IKZF3 have been described [21]. Since various T cell subsets and B cells are well-known to influence the nature and magnitude of the allergic immune response, future studies on IKZF3 could unravel its biological role in asthma pathogenesis. ZPBP2 is a protein coding gene implicated in gamete interaction during fertilization [22]. Diseases associated with ZPBP2 include osteogenesis imperfecta and this gene might not be directly involved in asthma biology.
Environmental exposures have a profound impact on the development and clinical course of childhood asthma. Environmental exposures commonly linked with asthma include tobacco smoke exposure, domestic and farm animals, respiratory viral infections, and occupational exposures [23]. Variants of the 17q21 locus may enhance the association between early respiratory infections and childhood asthma. Respiratory infections with human rhinovirus (HRV) in individuals with the risk genotype for rs7216389 SNP was associated with a >10- fold increase in odds ratio for childhood asthma. The rs7216389 genotype was also associated with increased transcript levels of ORMDL3 and GSDMB, but not IKZF3, in HRV-stimulated PBMCs [24]. The increased risk for early-onset asthma conferred by the 17q21 gene variants was also increased by early-life exposure to environmental tobacco smoke [4]. These studies highlight the interactions between 17q21 gene variants and common environmental risk factors, thus providing insights into the functional role of 17q21 locus in the pathophysiology of asthma.
Conclusion
In conclusion, several genetic epidemiological studies have consistently shown the association of 17q21 locus with asthma, thus highlighting the paramount importance of studying the biological roles of 17q21 genes. Recent experimental evidence has linked ORMDL3 to activation of ATF6α, altered calcium homeostasis, and sphingolipid synthesis. In addition, ORMDL3 transgenic mice exhibit increased airway remodeling and airway responsiveness suggesting an important role for ORMDL3 in asthma. At present the biological function(s) of the three other genes of this locus, including GSDMB, IKZF3 and ZPBP2 remains unexplored. As current asthma therapies mainly rely on symptomatic relief, identifying novel pathways that could be therapeutically manipulated based on a patient’s genotype is an important step towards developing a more personalized medicine approach for asthma therapies.
References
- Broide DH (2008) Immunologic and inflammatory mechanisms that drive asthma progression to remodeling. J Allergy Clin Immunol 121: 560-572.
- Moffatt MF, Gut IG, Demenais F, Strachan DP, Bouzigon E, et al. (2010) A large-scale, consortium-based genome wide association study of asthma. N Engl J Med 363: 1211-1221.
- Moffatt MF, Kabesch M, Liang L, Dixon AL, Strachan D, et al. (2007) Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448: 470-473.
- Bouzigon E, Corda E, Aschard H, Dizier MH, Boland A, et al. (2008) Effect of 17q21 variants and smoking exposure in early-onset asthma. N Engl J Med 359: 1985-1994.
- Zavbi M, Korosec P, Flezar M, Skrgat Kristan S, Marc Malovrh M, et al. (2016) Polymorphisms and haplotypes of the chromosome locus 17q12-17q21.1 contribute to adult asthma susceptibility in Slovenian patients. Hum Immunol 77: 527-534.
- Nieuwenhuis MA, Siedlinski M, van den Berge M, Granell R, Li X, et al. (2016) Combining Genome Wide Association Study and lung eQTL analysis provides evidence for novel genes associated with asthma. Allergy.
- Loss GJ, Depner M, Hose AJ, Genuneit J, Karvonen AM, et al. (2016) The Early Development of Wheeze. Environmental Determinants and Genetic Susceptibility at 17q21. Am J Respir Crit Care Med 193: 889-897.
- Galanter J, Choudhry S, Eng C, Nazario S, Rodriguez-Santana JR, et al. (2008) ORMDL3 gene is associated with asthma in three ethnically diverse populations. Am J Respir Crit Care Med 177: 1194-1200.
- Madore AM, Tremblay K, Hudson TJ, Laprise C (2008) Replication of an association between 17q21 SNPs and asthma in a French-Canadian familial collection. Hum Genet 123: 93-95.
- Li FX, Tan JY, Yang XX, Wu YS, Wu D, et al. (2012) Genetic variants on 17q21 are associated with asthma in a Han Chinese population. Genet Mol Res 11: 340-347.
- Hrdlickova B, Holla LI (2011) Relationship between the 17q21 locus and adult asthma in a Czech population. Hum Immunol 72: 921-925.
- Miller M, Tam AB, Cho JY, Doherty TA, Pham A, et al. (2012) ORMDL3 is an inducible lung epithelial gene regulating metallo proteases, chemokines, OAS and ATF6. Proc Natl Acad Sci USA 109: 16648-16653.
- Miller M, Rosenthal P, Beppu A, Mueller JL, Hoffman HM, et al. (2014) ORMDL3 transgenic mice have increased airway remodeling and airway responsiveness characteristic of asthma. J Immunol 192: 3475-3487.
- Levy BD (2013) Sphingolipids and Susceptibility to Asthma. N Engl J Med 369: 976-978.
- Ono JG, Worgall TS, Worgall S (2015) Airway reactivity and sphingolipids-implications for childhood asthma. Mol Cell Pediatr 2: 13.
- Ha SG, Ge XN, Bahaie NS, Kang BN, Rao A, et al. (2013) ORMDL3 promotes eosinophil trafficking and activation via regulation of integrins and CD48. Nat Commun 4: 2479-2479.
- Tamura M, Tanaka S, Fujii T, Aoki A, Komiyama H, et al. (2007) Members of a novel gene family, Gsdm, are expressed exclusively in the epithelium of the skin and gastrointestinal tract in a highly tissue-specific manner. Genomics 89: 618-629.
- Katoh M, Katoh M (2004) Evolutionary recombination hotspot around GSDML-GSDM locus is closely linked to the oncogenomic recombination hotspot around the PPP1R1B-ERBB2-GRB7 amplicon. Int J Oncol 24: 757-763.
- Hergueta-Redondo M, Sarrio D, Molina-Crespo A, Megias D, Mota A, et al. (2014) Gasdermin-B promotes invasion and metastasis in breast cancer cells. PLoS One 9: e90099.
- Rebollo A, Schmitt C (2003) Ikaros, Aiolos and Helios: Transcription regulators and lymphoid malignancies. Immunol Cell Biol 81: 171-175.
- Duhamel M, Arrouss I, Merle-Beral H, Rebollo A (2008) The Aiolos transcription factor is up-regulated in chronic lymphocytic leukemia. Blood 111: 3225-3228.
- Redgrove KA, Anderson AL, Dun MD, McLaughlin EA, O'Bryan MK, et al. (2011) Involvement of multimeric protein complexes in mediating the capacitation-dependent binding of human spermatozoa to homologous zonae pellucidae. Dev Biol 356: 460-474.
- Ober C, Yao TC (2011) The genetics of asthma and allergic disease: a 21st century perspective. Immunol Rev 242: 10-30.
- ÇalÃÆââ¬Å¾ÃâñÃÆââ¬Â¦Ãâà ¸kan M, Bochkov YA, Kreiner-Møller E, Bønnelykke K, Stein MM, et al. (2013) Rhinovirus Wheezing Illness and Genetic Risk of Childhood-Onset Asthma. N Engl J Med 368: 1398-1407.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences